Mammalia#
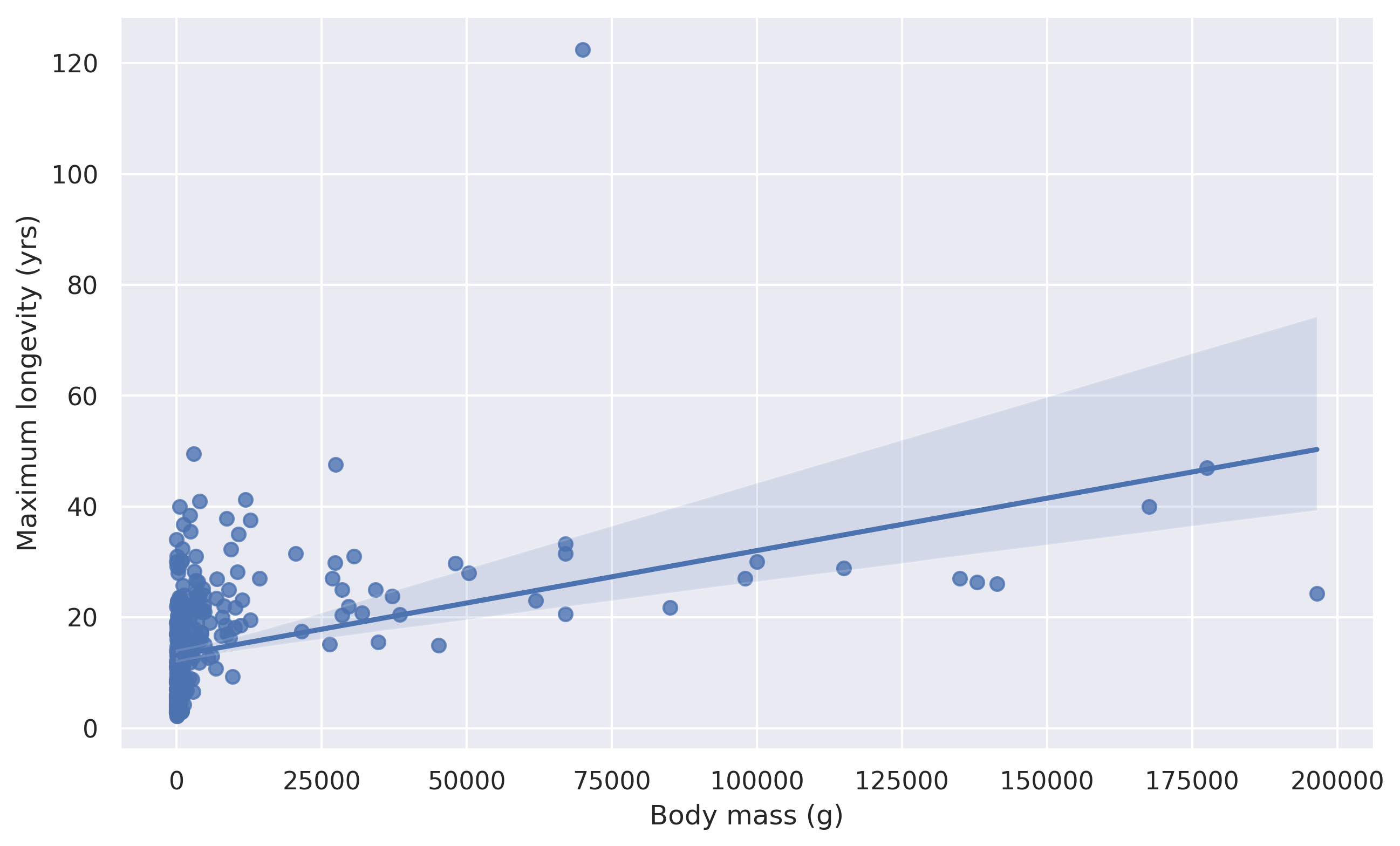
Generate a regression plot to visualize whether there is any linear relationship between body mass and maximum longevity of animals in the dataset. Only consider samples for the Mammalia class and a body mass of less than 200,000.
Importing libraries and packages#
1# Warnings
2import warnings
3
4# Mathematical operations and data manipulation
5import pandas as pd
6import numpy as np
7
8# Plotting
9import matplotlib.pyplot as plt
10import seaborn as sns
11
12sns.set()
13warnings.filterwarnings("ignore")
Set paths#
1# Path to datasets directory
2data_path = "./datasets"
3# Path to assets directory (for saving results to)
4assets_path = "./assets"
Loading dataset#
1dataset = pd.read_csv(f"{data_path}/anage_data.csv", index_col=0)
Exploring dataset#
1# Shape of the dataset
2print("Shape of the dataset: ", dataset.shape)
3# View
4dataset
Shape of the dataset: (4218, 29)
| HAGRID | Kingdom | Phylum | Class | Order | Family | Genus | Species | Common name | Female maturity (days) | ... | Growth rate (1/days) | Maximum longevity (yrs) | Specimen origin | Sample size | Data quality | IMR (per yr) | MRDT (yrs) | Metabolic rate (W) | Body mass (g) | Temperature (K) | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 3 | Animalia | Arthropoda | Branchiopoda | Diplostraca | Daphniidae | Daphnia | pulicaria | Daphnia | NaN | ... | NaN | 0.19 | unknown | medium | acceptable | NaN | NaN | NaN | NaN | NaN |
| 1 | 5 | Animalia | Arthropoda | Insecta | Diptera | Drosophilidae | Drosophila | melanogaster | Fruit fly | 7.0 | ... | NaN | 0.30 | captivity | large | acceptable | 0.05 | 0.04 | NaN | NaN | NaN |
| 2 | 6 | Animalia | Arthropoda | Insecta | Hymenoptera | Apidae | Apis | mellifera | Honey bee | NaN | ... | NaN | 8.00 | unknown | medium | acceptable | NaN | NaN | NaN | NaN | NaN |
| 3 | 8 | Animalia | Arthropoda | Insecta | Hymenoptera | Formicidae | Cardiocondyla | obscurior | Cardiocondyla obscurior | NaN | ... | NaN | 0.50 | captivity | medium | acceptable | NaN | NaN | NaN | NaN | NaN |
| 4 | 9 | Animalia | Arthropoda | Insecta | Hymenoptera | Formicidae | Lasius | niger | Black garden ant | NaN | ... | NaN | 28.00 | unknown | medium | acceptable | NaN | NaN | NaN | NaN | NaN |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
| 4214 | 4239 | Animalia | Porifera | Hexactinellida | Lyssacinosida | Rossellidae | Scolymastra | joubini | Hexactinellid sponge | NaN | ... | NaN | 15000.00 | wild | medium | questionable | NaN | NaN | NaN | NaN | NaN |
| 4215 | 4241 | Plantae | Pinophyta | Pinopsida | Pinales | Pinaceae | Pinus | longaeva | Great Basin bristlecone pine | NaN | ... | NaN | 5062.00 | wild | medium | acceptable | NaN | 999.00 | NaN | NaN | NaN |
| 4216 | 4242 | Fungi | Ascomycota | Saccharomycetes | Saccharomycetales | Saccharomycetaceae | Saccharomyces | cerevisiae | Baker's yeast | NaN | ... | NaN | 0.04 | captivity | large | acceptable | NaN | NaN | NaN | NaN | NaN |
| 4217 | 4243 | Fungi | Ascomycota | Schizosaccharomycetes | Schizosaccharomycetales | Schizosaccharomycetaceae | Schizosaccharomyces | pombe | Fission yeast | NaN | ... | NaN | NaN | unknown | small | low | NaN | NaN | NaN | NaN | NaN |
| 4218 | 4244 | Fungi | Ascomycota | Sordariomycetes | Sordariales | Lasiosphaeriaceae | Podospora | anserina | Filamentous fungus | NaN | ... | NaN | NaN | unknown | small | low | NaN | NaN | NaN | NaN | NaN |
4218 rows × 29 columns
Preprocessing#
1longevity = "Maximum longevity (yrs)"
2mass = "Body mass (g)"
3data = dataset[dataset["Class"] == "Mammalia"]
4data = data[
5 np.isfinite(data[longevity])
6 & np.isfinite(data[mass])
7 & (data[mass] < 200000)
8]
Visualisation#
1# Create regression plot
2plt.figure(figsize=(10, 6), dpi=300)
3# Create scatter plot
4sns.regplot(mass, longevity, data=data)
5# Show plot
6plt.show()