Exploratory data analysis#
Importing libraries and packages#
1# Mathematical operations and data manipulation
2import numpy as np
3import pandas as pd
4
5# Visualisation
6import seaborn as sns
7import matplotlib.pyplot as plt
8
9# Warnings
10import warnings
11
12warnings.filterwarnings("ignore")
13
14%matplotlib inline
Set paths#
1# Path to datasets directory
2data_path = "./datasets"
3# Path to assets directory (for saving results to)
4assets_path = "./assets"
Loading dataset#
1# load data
2dataset = pd.read_csv(f"{data_path}/preprocessed_heart.csv")
3dataset.head().T
| 0 | 1 | 2 | 3 | 4 | |
|---|---|---|---|---|---|
| age | 63.0 | 37.0 | 41.0 | 56.0 | 57.0 |
| sex | 1.0 | 1.0 | 0.0 | 1.0 | 0.0 |
| chest_pain | 3.0 | 2.0 | 1.0 | 1.0 | 0.0 |
| rest_bp | 145.0 | 130.0 | 130.0 | 120.0 | 120.0 |
| chol | 233.0 | 250.0 | 204.0 | 236.0 | 354.0 |
| fast_bld_sugar | 1.0 | 0.0 | 0.0 | 0.0 | 0.0 |
| rest_ecg | 0.0 | 1.0 | 0.0 | 1.0 | 1.0 |
| max_hr | 150.0 | 187.0 | 172.0 | 178.0 | 163.0 |
| ex_angina | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 |
| st_depr | 2.3 | 3.5 | 1.4 | 0.8 | 0.6 |
| slope | 0.0 | 0.0 | 2.0 | 2.0 | 2.0 |
| colored_vessels | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
| thalassemia | 1.0 | 2.0 | 2.0 | 2.0 | 2.0 |
| target | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 |
Plotting distributions and relationships#
Plotting the Distributions and Relationships Between Specific Features#
1sns.set(
2 palette="pastel",
3 rc={
4 "figure.figsize": (16, 10),
5 "axes.titlesize": 18,
6 "axes.labelsize": 16,
7 "xtick.labelsize": 14,
8 "ytick.labelsize": 16,
9 },
10)
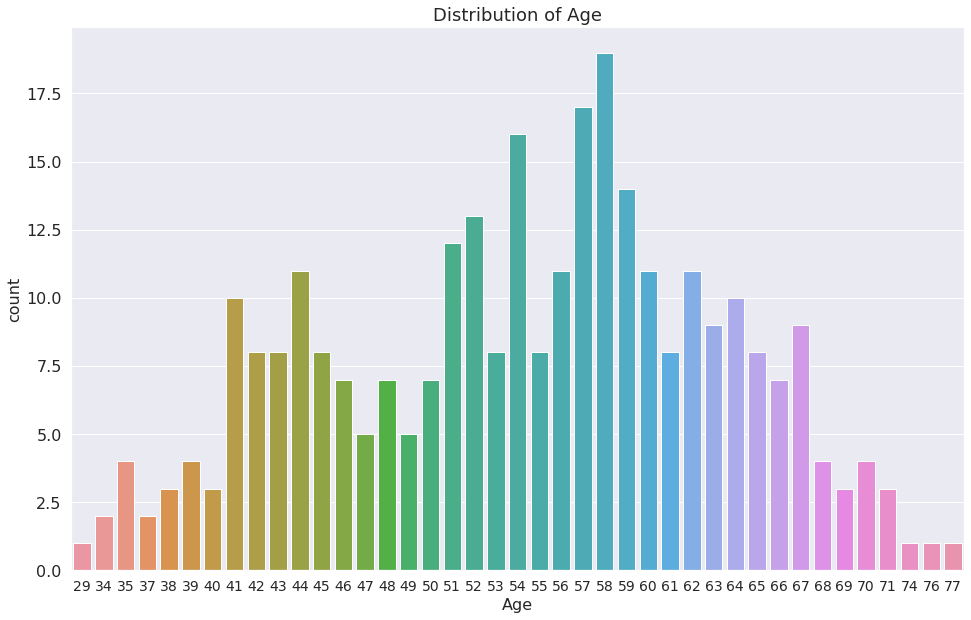
1g = sns.countplot(x="age", data=dataset)
2g.set_title("Distribution of Age")
3plt.xlabel("Age")
Text(0.5, 0, 'Age')
1print(dataset["target"].value_counts())
2print()
3print(dataset["target"].value_counts(normalize=True))
1 165
0 138
Name: target, dtype: int64
1 0.544554
0 0.455446
Name: target, dtype: float64
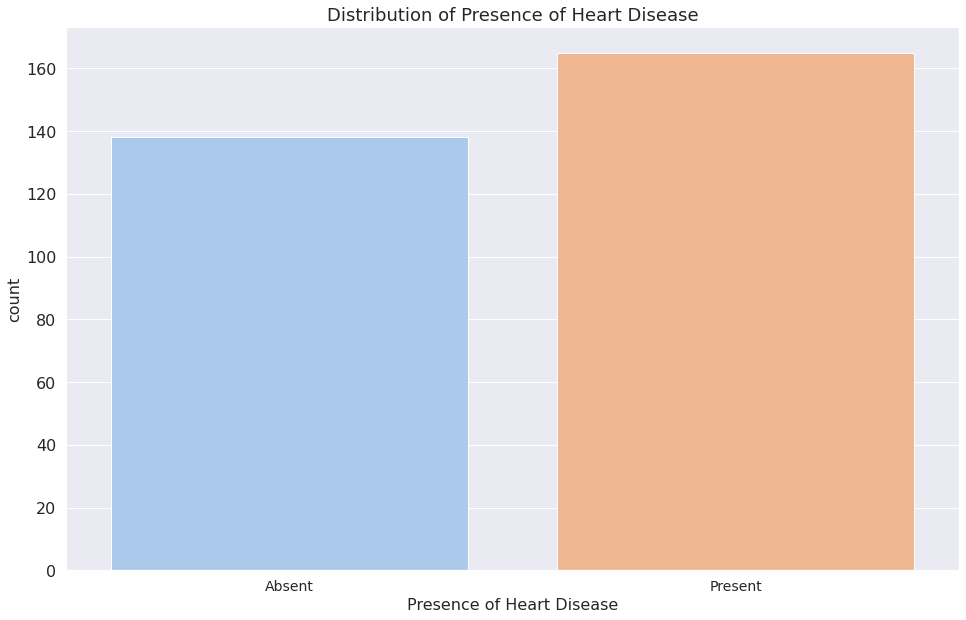
1a = sns.countplot(x="target", data=dataset)
2a.set_title("Distribution of Presence of Heart Disease")
3a.set_xticklabels(["Absent", "Present"])
4plt.xlabel("Presence of Heart Disease")
5plt.show()
1print(dataset["sex"].value_counts())
2print()
3print(dataset["sex"].value_counts(normalize=True))
1 207
0 96
Name: sex, dtype: int64
1 0.683168
0 0.316832
Name: sex, dtype: float64
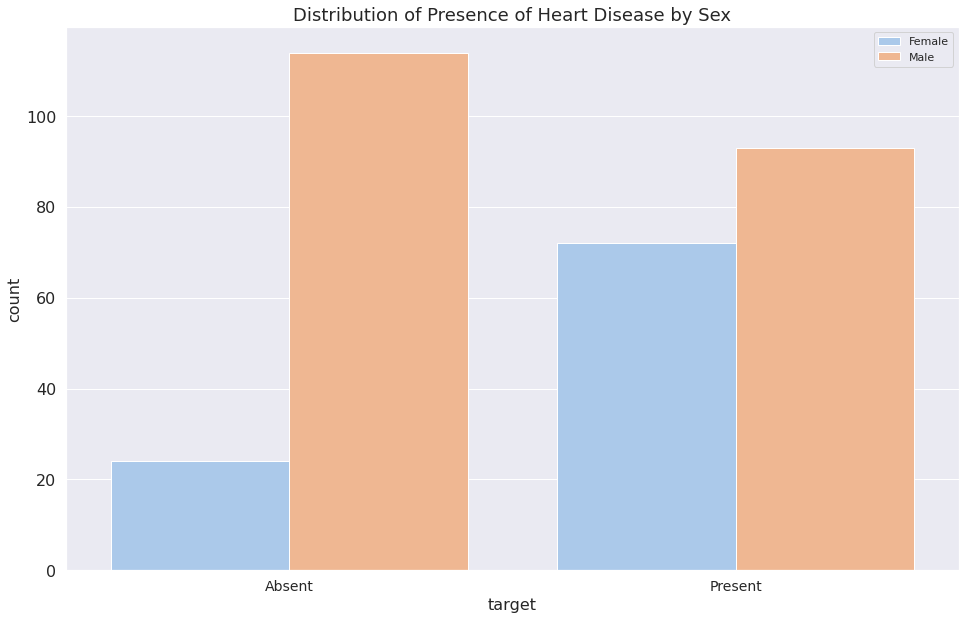
1b = sns.countplot(x="target", data=dataset, hue="sex")
2plt.legend(["Female", "Male"])
3b.set_title("Distribution of Presence of Heart Disease by Sex")
4b.set_xticklabels(["Absent", "Present"])
5plt.show()
Plotting Distributions and Relationships between Columns with Respect to the Target Column#
1print(dataset["chest_pain"].value_counts())
2print()
3print(dataset["chest_pain"].value_counts(normalize=True))
0 143
2 87
1 50
3 23
Name: chest_pain, dtype: int64
0 0.471947
2 0.287129
1 0.165017
3 0.075908
Name: chest_pain, dtype: float64
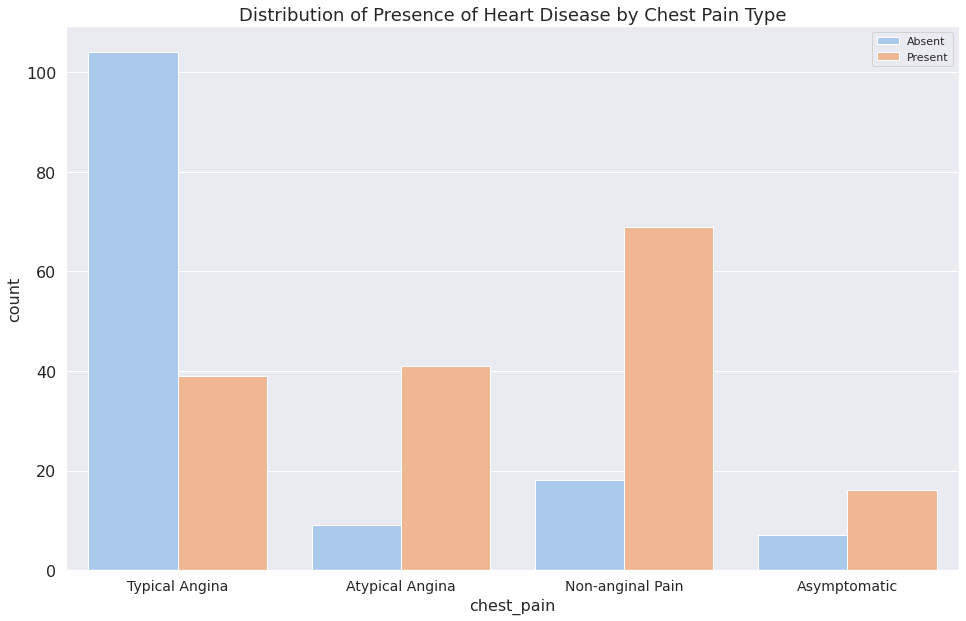
1c = sns.countplot(x="chest_pain", data=dataset, hue="target")
2plt.legend(["Absent", "Present"])
3c.set_title("Distribution of Presence of Heart Disease by Chest Pain Type")
4c.set_xticklabels(
5 ["Typical Angina", "Atypical Angina", "Non-anginal Pain", "Asymptomatic"]
6)
7plt.show()
1print(dataset["colored_vessels"].value_counts())
2print()
3print(dataset["colored_vessels"].value_counts(normalize=True))
0 175
1 65
2 38
3 20
4 5
Name: colored_vessels, dtype: int64
0 0.577558
1 0.214521
2 0.125413
3 0.066007
4 0.016502
Name: colored_vessels, dtype: float64
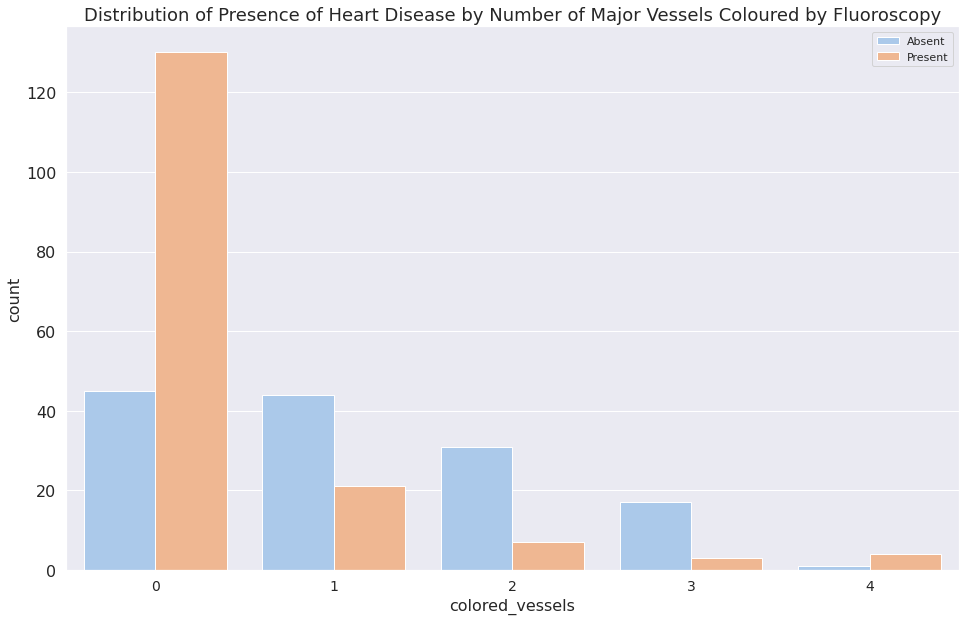
1d = sns.countplot(x="colored_vessels", data=dataset, hue="target")
2plt.legend(["Absent", "Present"])
3d.set_title(
4 "Distribution of Presence of Heart Disease by Number of Major "
5 "Vessels Coloured by Fluoroscopy"
6)
7plt.show()
1print(dataset["slope"].value_counts())
2print()
3print(dataset["slope"].value_counts(normalize=True))
2 142
1 140
0 21
Name: slope, dtype: int64
2 0.468647
1 0.462046
0 0.069307
Name: slope, dtype: float64
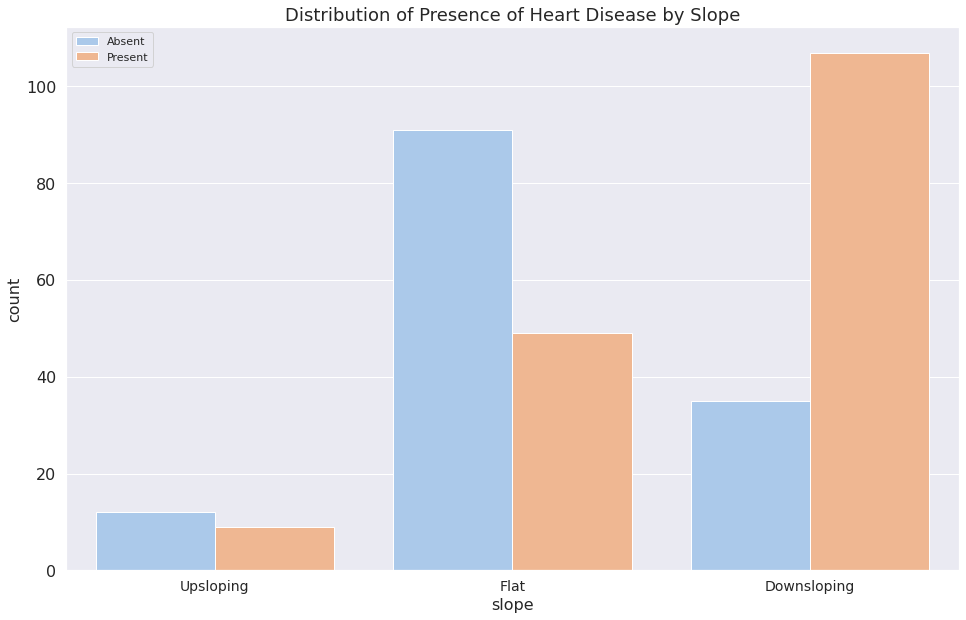
1f = sns.countplot(x="slope", data=dataset, hue="target")
2plt.legend(["Absent", "Present"])
3f.set_title("Distribution of Presence of Heart Disease by Slope")
4f.set_xticklabels(["Upsloping", "Flat", "Downsloping"])
5plt.show()
Plotting the Relationship between the Presence of Heart Disease and Maximum Recorded Heart Rate#
1# Plotting the Relationship between the Presence of Heart Disease and
2# Maximum Recorded Heart Rate
3sns.set(
4 style="whitegrid",
5 palette="colorblind",
6 rc={
7 "figure.figsize": (12, 8),
8 "axes.titlesize": 18,
9 "axes.labelsize": 16,
10 "xtick.labelsize": 16,
11 "ytick.labelsize": 16,
12 },
13)
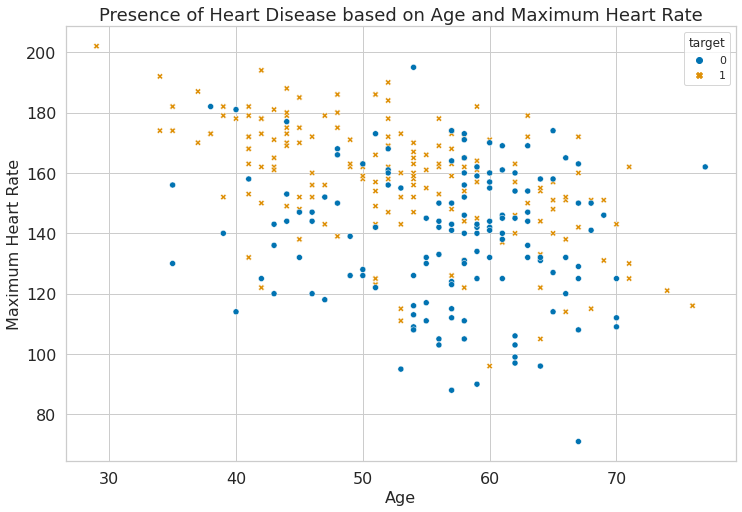
1f = sns.scatterplot(
2 x="age", y="max_hr", hue="target", style="target", data=dataset
3)
4f.set_title("Presence of Heart Disease based on Age and Maximum Heart Rate")
5plt.xlabel("Age")
6plt.ylabel("Maximum Heart Rate")
Text(0, 0.5, 'Maximum Heart Rate')
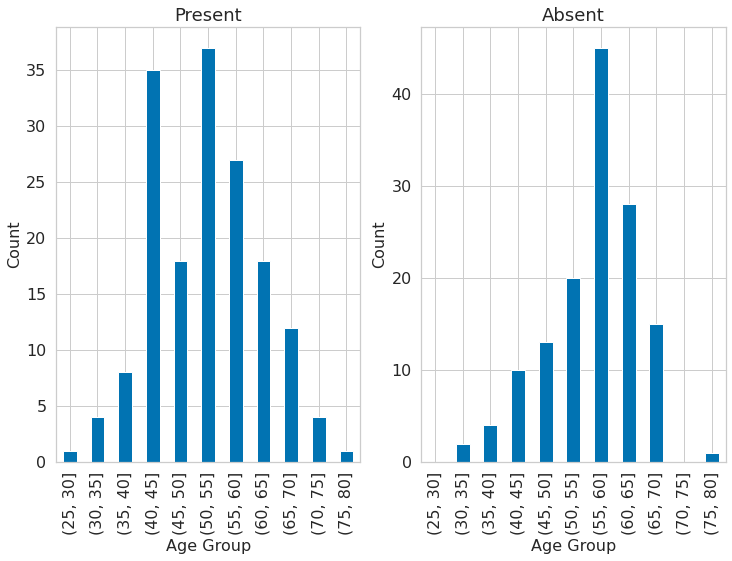
1dataset["age_category"] = pd.cut(dataset.age, bins=list(np.arange(25, 85, 5)))
2
3plt.subplot(121)
4dataset[dataset.target == 1].groupby("age_category")["age"].count().plot(
5 kind="bar"
6)
7plt.title("Present")
8plt.xlabel("Age Group")
9plt.ylabel("Count")
10plt.subplot(122)
11dataset[dataset.target == 0].groupby("age_category")["age"].count().plot(
12 kind="bar"
13)
14plt.title("Absent")
15plt.xlabel("Age Group")
16plt.ylabel("Count")
Text(0, 0.5, 'Count')
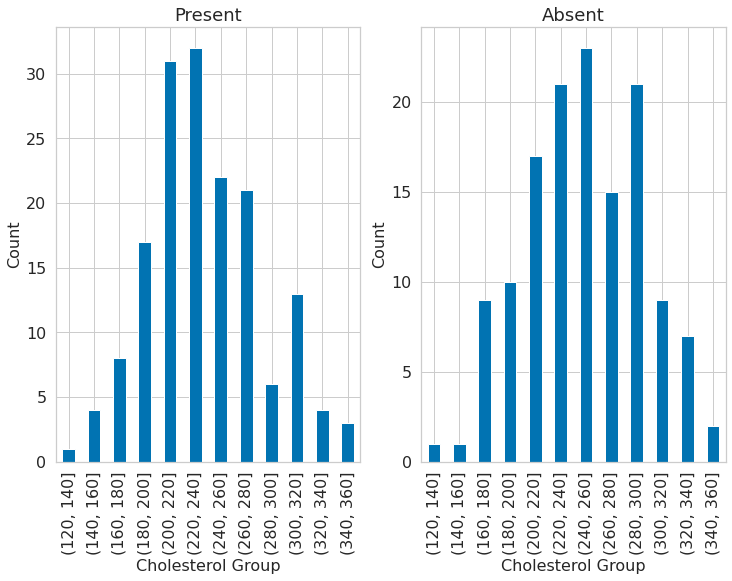
Plotting the Relationship between the Presence of Heart Disease and the Cholesterol Column#
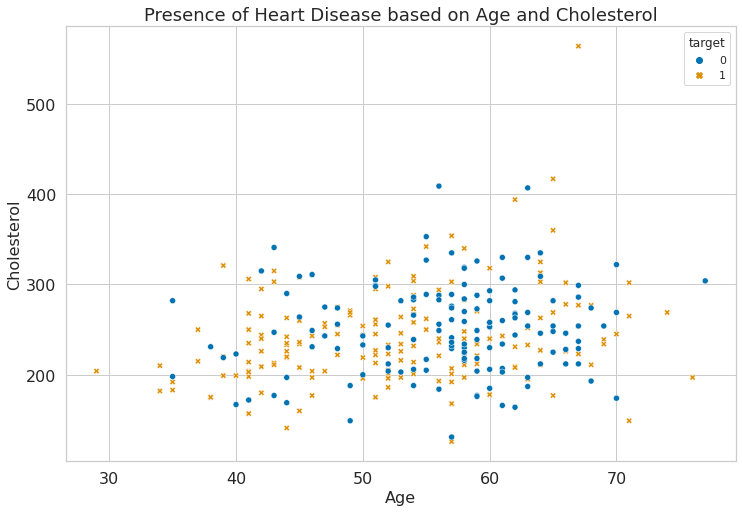
1g = sns.scatterplot(
2 x="age", y="chol", hue="target", style="target", data=dataset
3)
4g.set_title("Presence of Heart Disease based on Age and Cholesterol")
5plt.xlabel("Age")
6plt.ylabel("Cholesterol")
Text(0, 0.5, 'Cholesterol')
1dataset["chol_cat"] = pd.cut(dataset.chol, bins=list(np.arange(120, 380, 20)))
2dataset["chol_cat"] = pd.cut(dataset.chol, bins=list(np.arange(120, 380, 20)))
3
4plt.subplot(121)
5dataset[dataset.target == 1].groupby("chol_cat")["chol"].count().plot(
6 kind="bar"
7)
8plt.title("Present")
9plt.xlabel("Cholesterol Group")
10plt.ylabel("Count")
11
12plt.subplot(122)
13dataset[dataset.target == 0].groupby("chol_cat")["chol"].count().plot(
14 kind="bar"
15)
16plt.title("Absent")
17plt.xlabel("Cholesterol Group")
18plt.ylabel("Count")
Text(0, 0.5, 'Count')